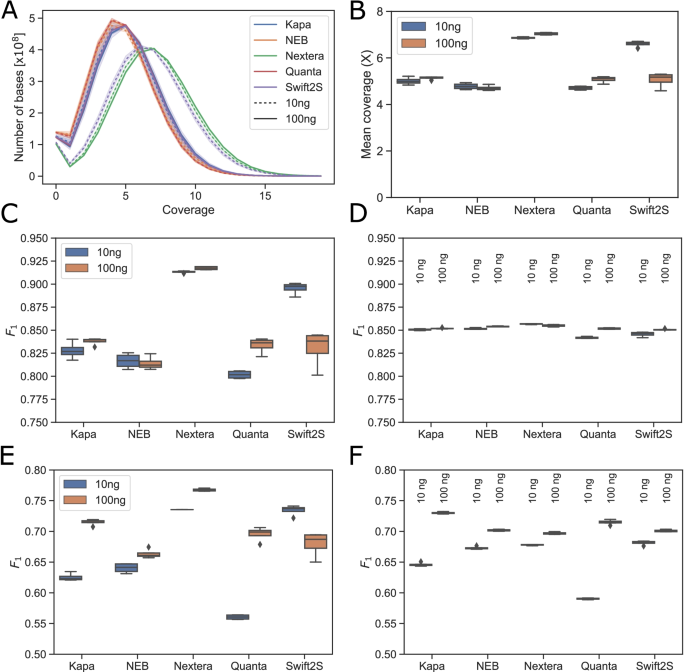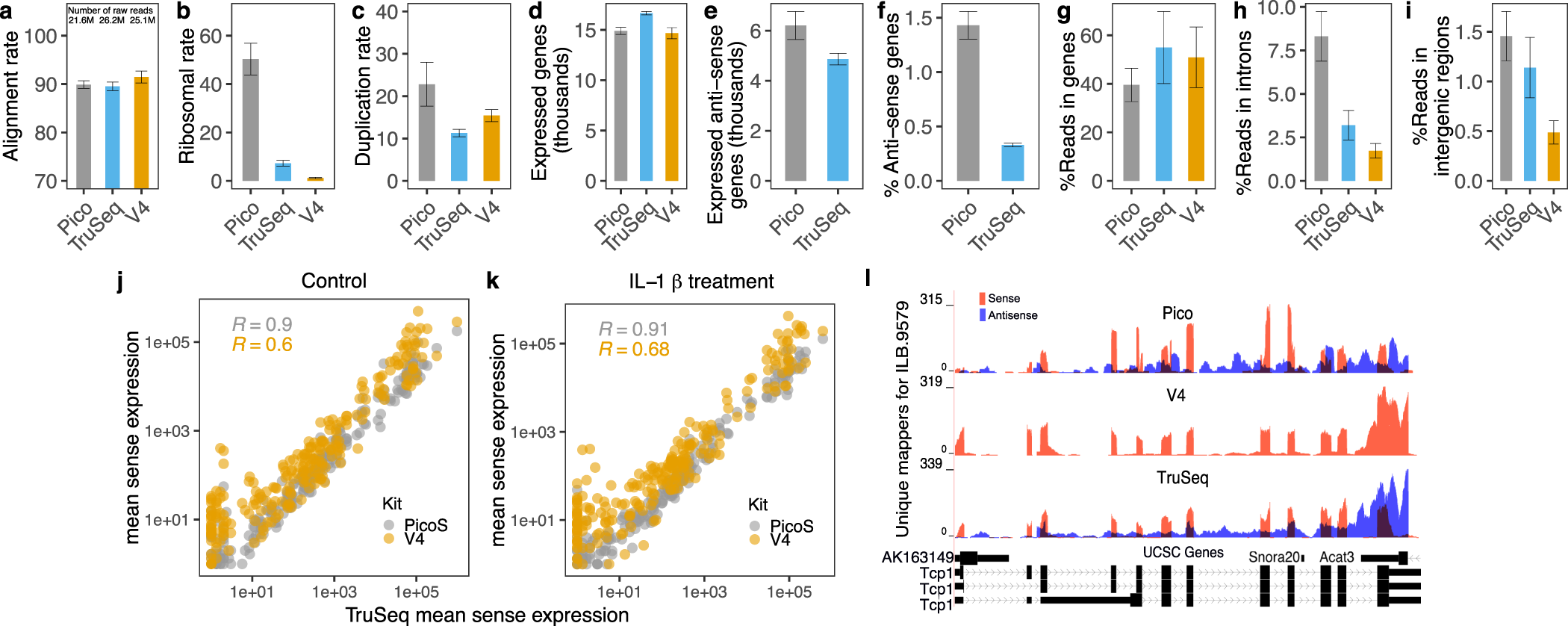
Comparative evaluation of RNA-Seq library preparation methods for strand-specificity and low input | Scientific Reports

Evaluation of a high-throughput, cost-effective Illumina library preparation kit | Scientific Reports

Best practices to minimize rRNA contamination in TruSeq Stranded Total RNA libraries - Illumina Knowledge
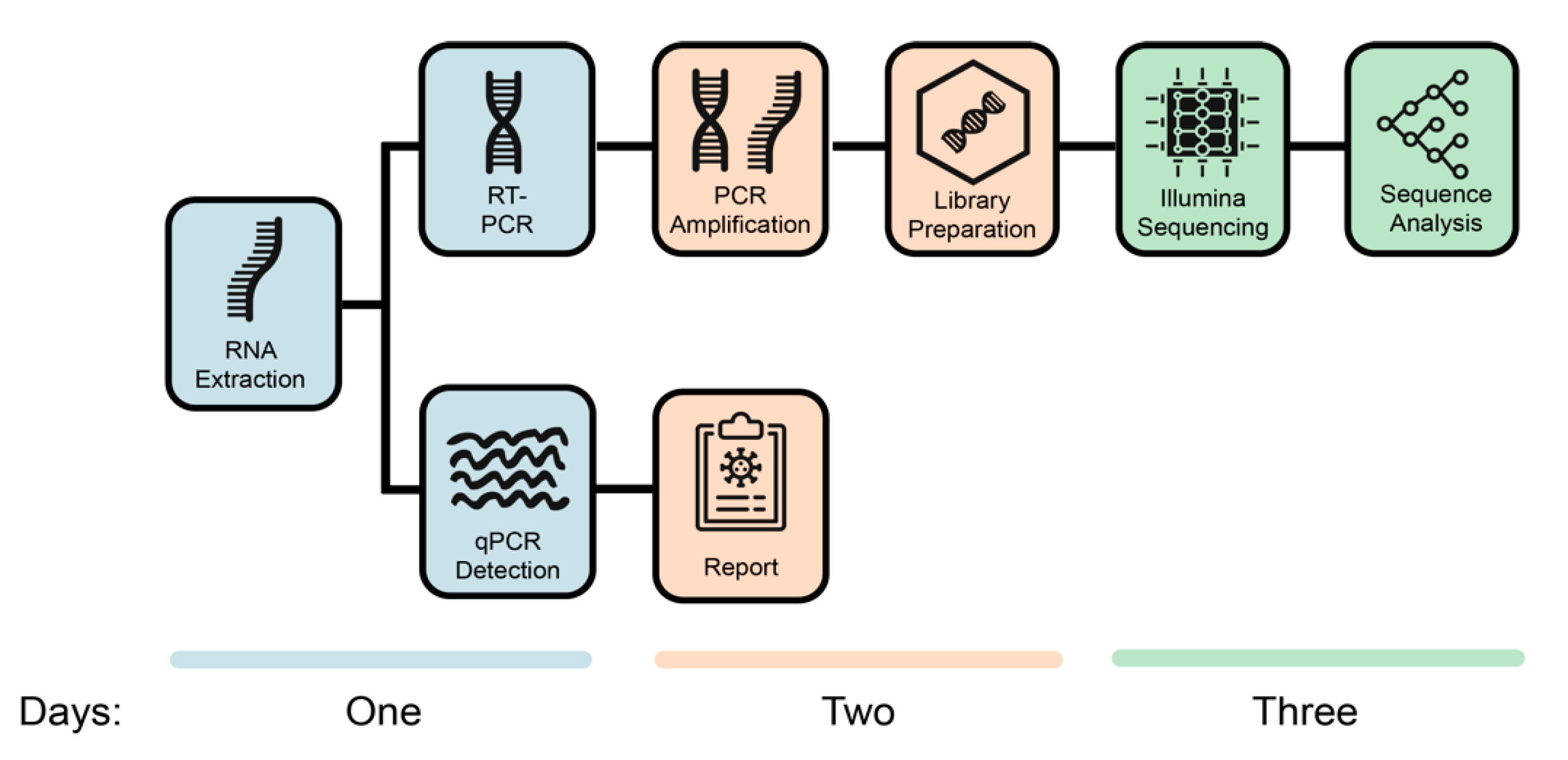
Genes | Free Full-Text | Whole Genome Sequencing of SARS-CoV-2: Adapting Illumina Protocols for Quick and Accurate Outbreak Investigation during a Pandemic

Bubble products in sequencing libraries: causes, identification, and workflow recommendations - Illumina Knowledge